Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 6(5); 2015 > Article
-
Brief Report
Occurrence of Norovirus GII.4 Sydney Variant-related Outbreaks in Korea - Sunyoung Junga, Bo-Mi Hwanga, Hyun Ju Jeonga, Gyung Tae Chunga, Cheon-Kwon Yooa, Yeon-Ho Kangb, Deog-Yong Leea
-
Osong Public Health and Research Perspectives 2015;6(5):322-326.
DOI: https://doi.org/10.1016/j.phrp.2015.10.004
Published online: October 22, 2015
aDivision of Enteric Diseases, Center for Infectious Diseases, Korea National Institute of Health, KCDC, Osong, Korea
bDivision of Bio-Safety Evaluation and Control, Korea National Institute of Health, KCDC, Osong, Korea
- ∗Corresponding author. leedy0610@korea.kr
• Received: February 5, 2015 • Revised: October 2, 2015 • Accepted: October 2, 2015
Copyright © 2015 Korea Centers for Disease Control and Prevention. Published by Elsevier Korea LLC. All rights reserved.
Abstract
- Human noroviruses are major causative agents of food and waterborne outbreaks of nonbacterial acute gastroenteritis. In this study, we report the epidemiological features of three outbreak cases of norovirus in Korea, and we describe the clinical symptoms and distribution of the causative genotypes. The incidence rates of the three outbreaks were 16.24% (326/2,007), 4.1% (27/656), and 16.8% (36/214), respectively. The patients in these three outbreaks were affected by acute gastroenteritis. These schools were provided unheated food from the same manufacturing company. Two genotypes (GII.3 and GII.4) of the norovirus were detected in these cases. Among them, major causative strains of GII.4 (Hu-jeju-47-2007KR-like) were identified in patients, food handlers, and groundwater from the manufacturing company of the unheated food. In the GII.4 (Hu-jeju-47-2007KR-like) strain of the norovirus, the nucleotide sequences were identical and identified as the GII.4 Sydney variant. Our data suggests that the combined epidemiological and laboratory results were closely related, and the causative pathogen was the GII.4 Sydney variant strain from contaminated groundwater.
- The authors have nothing to declare.
Conflicts of interest
-
Acknowledgements
- This study was supported by the National Norovirus Surveillance System (4851-304-210) and conducted by the Korea National Institute Health and Seoul and Gyungbuk Institute of Health and Environment Research.
Acknowledgments
-
This is an open-access article distributed under the terms of the Creative Commons Attribution-NonCommercial-No Derivative Works License (http://creativecommons.org/licenses/by-nc-nd/4.0) which permits non-commercial use, distribution, and reproduction in any medium, provided the original author and source are credited.
Article information
- 1. Rockx B., De Wit M., Vennema H.. Natural history of human calicivirus infection: a prospective cohort study. Clin Infect Dis 35(3). 2002 Aug;246−253. PMID: 12115089.ArticlePubMed
- 2. Maunula L., Miettinen I.T., von Bonsdorff C.H.. Norovirus outbreaks from drinking water. Emerg Infect Dis 11(11). 2005 Nov;1716−1721. PMID: 16318723.ArticlePubMed
- 3. Hewitt J., Bell D., Simmons G.C.. Gastroenteritis outbreak caused by waterborne norovirus at a New Zealand ski resort. Appl Environ Microbiol 73(24). 2007 Dec;7853−7857. PMID: 17965205.ArticlePubMed
- 4. Belliot G., Kamel A.H., Estienney M.. Evidence of emergence of new GGII.4 norovirus variants from gastroenteritis outbreak survey in France during the 2007-to-2008 and 2008-to-2009 winter seasons. J Clin Microbiol 48(3). 2010 Mar;994−998. PMID: 20042616.ArticlePubMed
- 5. Vega E., Barclay L., Gregoricus N.. Novel surveillance network for norovirus gastroenteritis outbreaks, United States. Emerg Infect Dis 17(8). 2011 Aug;1389−1395. PMID: 21801614.ArticlePubMed
- 6. van Beek J., Ambert-Balay K., Botteldoorn N.. Indications for worldwide increased norovirus activity associated with emergence of a new variant of genotype II.4, late 2012. Euro Surveill 18(1). 2013 Jan;8−9. PMID: 23305715.PubMed
- 7. Felsenstein J.. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39(4). 1985 Jul;783−791.ArticlePubMed
- 8. Saitou N., Nei M.. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4). 1987 Jul;406−425. PMID: 3447015.PubMed
- 9. Fonager J., Hindbæk L.S., Fischer T.K.. Rapid emergence and antigenic diversification of the norovirus 2012 Sydney variant in Denmark, October to December, 2012. Euro Surveill 18(9). 2013 Feb;28;1−4. PMID: 23449181.Article
- 10. Polkowska A., Ronnqvist M., Lepisto O.. Outbreak of gastroenteritis caused by norovirus GII.4 Sydney variant after a wedding reception at a resort/activity centre, Finland, August 2012. Epidemiol Infect 142(9). 2014 Sep;1877−1883. PMID: 24229743.ArticlePubMed
- 11. Wong T.H., Dearlove B.L., Hedge J.. Whole genome sequencing and de novo assembly identifies Sydney-like variant noroviruses and recombinants during the winter 2012/2013 outbreak in England. Virol J 10:2013 Nov;335PMID: 24220146.ArticlePubMed
References
Figure 1Phylogenetic analysis of the norovirus detected from patients, food handlers, and environmental samples (unheated foods, supplemental water, and groundwater). The phylogenetic tree was constructed with the neighbor-joining method with norovirus partial capsid region (312-314bp). The numbers in the branches indicate the bootstrap values.
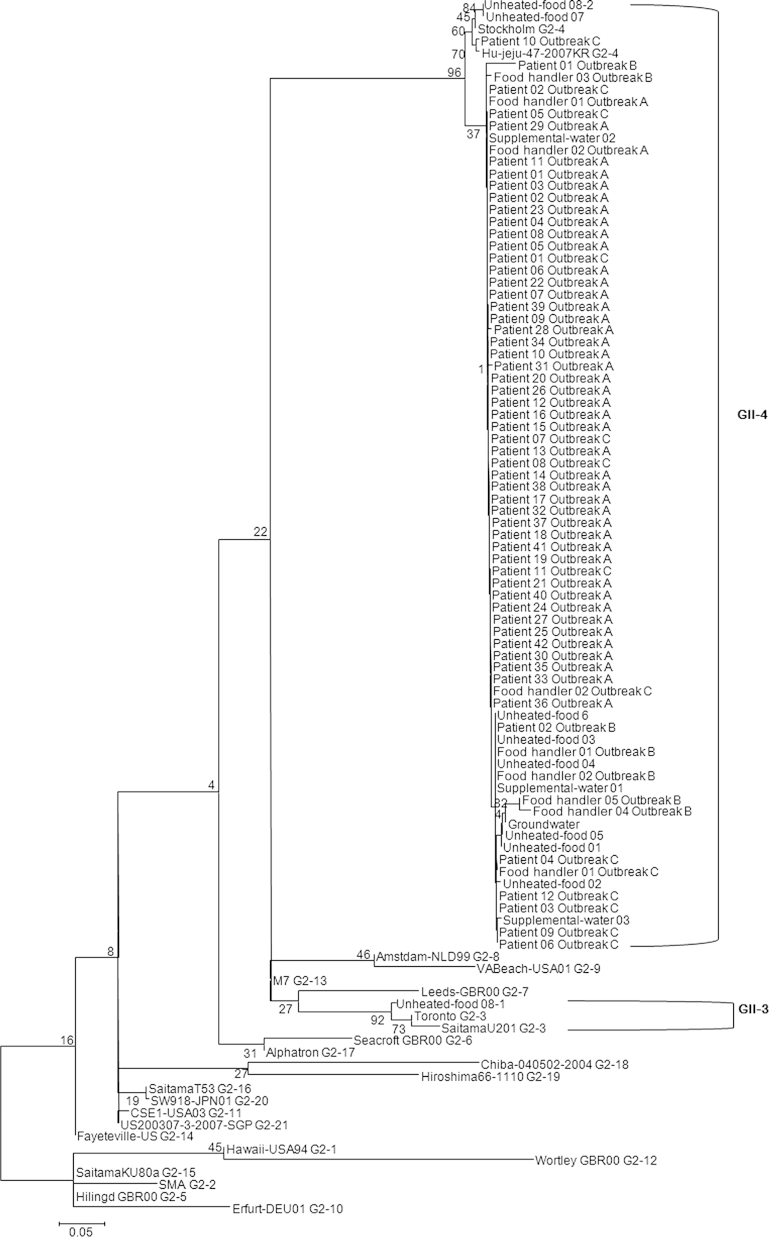

Table 1Comparison of epidemiological and laboratory characteristics.
Table 2Oligo nucleotide sequences of primers used in this study.
Figure & Data
References
Citations
Citations to this article as recorded by 

- A systematic review and meta-analysis indicates a substantial burden of human noroviruses in shellfish worldwide, with GII.4 and GII.2 being the predominant genotypes
Yijing Li, Liang Xue, Junshan Gao, Weicheng Cai, Zilei Zhang, Luobing Meng, Shuidi Miao, Xiaojing Hong, Mingfang Xu, Qingping Wu, Jumei Zhang
Food Microbiology.2023; 109: 104140. CrossRef - Molecular epidemiology of norovirus infections in children with acute gastroenteritis in 2017–2019 in Tianjin, China
Yulian Fang, Yanzhi Zhang, Hong Wang, Ouyan Shi, Wei Wang, Mengzhu Hou, Lu Wang, Jinying Wu, Yu Zhao
Journal of Medical Virology.2022; 94(2): 616. CrossRef - Assessment of potential infectivity of human norovirus in the traditional Korean salted clam product “Jogaejeotgal” by floating electrode-dielectric barrier discharge plasma
Eun Bi Jeon, Man-Seok Choi, Ji Yoon Kim, Eun Ha Choi, Jun Sup Lim, Jinsung Choi, Kwang Soo Ha, Ji Young Kwon, Sang Hyeon Jeong, Shin Young Park
Food Research International.2021; 141: 110107. CrossRef - Characterizing the effects of thermal treatment on human norovirus GII.4 viability using propidium monoazide combined with RT-qPCR and quality assessments in mussels
Eun Bi Jeon, Man-Seok Choi, Ji Yoon Kim, Kwang Soo Ha, Ji Young Kwon, Sung Hyeon Jeong, Hee Jung Lee, Yeoun Joong Jung, Ji-Hyoung Ha, Shin Young Park
Food Control.2020; 109: 106954. CrossRef - Molecular epidemiology of genogroup II norovirus infections in acute gastroenteritis patients during 2014–2016 in Pudong New Area, Shanghai, China
Caoyi Xue, Lifeng Pan, Weiping Zhu, Yuanping Wang, Huiqin Fu, Chang Cui, Lan Lu, Sun Qiao, Biao Xu
Gut Pathogens.2018;[Epub] CrossRef - Review: Epidemiological evidence of groundwater contribution to global enteric disease, 1948–2015
Heather M. Murphy, Morgan D. Prioleau, Mark A. Borchardt, Paul D. Hynds
Hydrogeology Journal.2017; 25(4): 981. CrossRef - Change in Concentrations of Human Norovirus and Male-Specific Coliphage under Various Temperatures, Salinities, and pH Levels in Seawater
Poong Ho Kim, Yong Soo Park, Kunbawui Park, Ji Young Kwon, Hong Sik Yu, Hee Jung Lee, Ji Hoe Kim, Tae Seek Lee
Korean Journal of Fisheries and Aquatic Sciences.2016; 49(4): 454. CrossRef - Norovirus outbreaks occurred in different settings in the Republic of Korea
Hae-Wol Cho, Chaeshin Chu
Osong Public Health and Research Perspectives.2015; 6(5): 281. CrossRef


 PubReader
PubReader Cite
Cite
