Articles
- Page Path
- HOME > Osong Public Health Res Perspect > Volume 11(3); 2020 > Article
-
Original Article
Detection and Isolation of SARS-CoV-2 in Serum, Urine, and Stool Specimens of COVID-19 Patients from the Republic of Korea -
Jeong-Min Kima
 , Heui Man Kima
, Heui Man Kima , Eun Jung Leea
, Eun Jung Leea , Hye Jun Joa
, Hye Jun Joa , Youngsil Yoona
, Youngsil Yoona , Nam-Joo Leea
, Nam-Joo Leea , Junseock Sona
, Junseock Sona , Ye-Ji Leea
, Ye-Ji Leea , Mi Seon Kima
, Mi Seon Kima , Yong-Pyo Leea
, Yong-Pyo Leea , Su-Jin Chaea
, Su-Jin Chaea , Kye Ryeong Parka
, Kye Ryeong Parka , Seung-Rye Choa
, Seung-Rye Choa , Sehee Parka
, Sehee Parka , Su Jin Kima
, Su Jin Kima , Eunbyeol Wanga
, Eunbyeol Wanga , SangHee Wooa
, SangHee Wooa , Aram Lima
, Aram Lima , Su-Jin Parka
, Su-Jin Parka , JunHyeong Janga
, JunHyeong Janga , Yoon-Seok Chunga
, Yoon-Seok Chunga , Bum Sik Chinb
, Bum Sik Chinb , Jin-Soo Leec
, Jin-Soo Leec , Duko Limd
, Duko Limd , Myung-Guk Hana
, Myung-Guk Hana , Cheon Kwon Yooe
, Cheon Kwon Yooe
-
Osong Public Health and Research Perspectives 2020;11(3):112-117.
DOI: https://doi.org/10.24171/j.phrp.2020.11.3.02
Published online: May 8, 2020
aDivision of Viral Diseases, Center for Laboratory Control of Infectious Diseases, Korea Centers for Disease Control and Prevention, Cheongju, Korea
bDivision of Infectious Diseases, Department of Internal Medicine, National Medical Center, Seoul, Korea
cDivision of Infectious Diseases and Infection Control Unit, Department of Internal Medicine, Inha University School of Medicine, Incheon, Korea
dDivision of Emergency Medicine, Gyeonggi Provincial Medical Center Ansung Hospital, Ansung, Korea
eCenter for Laboratory Control of Infectious Diseases, Korea Centers for Disease Control and Prevention, Cheongju, Korea
- *Corresponding author: Cheon Kwon Yoo Center for Laboratory Control of Infectious Diseases, Korea Centers for Disease Control and Prevention, Cheongju, Korea E-mail: ckyoo@korea.kr
Copyright © 2020 Korea Centers for Disease Control & Prevention
This is an open access article under the CC BYNC-ND license (http://creativecommons.org/licenses/by-nc/4.0/).
Abstract
-
Objectives
- Coronavirus Disease-19 (COVID-19) is a respiratory infection characterized by the main symptoms of pneumonia and fever. It is caused by the novel coronavirus severe acute respiratory syndrome Coronavirus-2 (SARS-CoV-2), which is known to spread via respiratory droplets. We aimed to determine the rate and likelihood of SARS-CoV-2 transmission from COVID-19 patients through non-respiratory routes.
-
Methods
- Serum, urine, and stool samples were collected from 74 hospitalized patients diagnosed with COVID-19 based on the detection of SARS-CoV-2 in respiratory samples. The SARS-CoV-2 RNA genome was extracted from each specimen and real-time reverse transcription polymerase chain reaction performed. CaCo-2 cells were inoculated with the specimens containing the SARS-COV-2 genome, and subcultured for virus isolation. After culturing, viral replication in the cell supernatant was assessed.
-
Results
- Of the samples collected from 74 COVID-19 patients, SARS-CoV-2 was detected in 15 serum, urine, or stool samples. The virus detection rate in the serum, urine, and stool samples were 2.8% (9/323), 0.8% (2/247), and 10.1% (13/129), and the mean viral load was 1,210 ± 1,861, 79 ± 30, and 3,176 ± 7,208 copy/µL, respectively. However, the SARS-CoV-2 was not isolated by the culture method from the samples that tested positive for the SARS-CoV-2 gene.
-
Conclusion
- While the virus remained detectable in the respiratory samples of COVID-19 patients for several days after hospitalization, its detection in the serum, urine, and stool samples was intermittent. Since the virus could not be isolated from the SARS-COV-2-positive samples, the risk of viral transmission via stool and urine is expected to be low.
- The Coronavirus Disease-19 (COVID-19) leads to the development of Severe Acute Respiratory Syndrome (SARS), with a fever, cough, sore throat, myalgia, and headache as the most common clinical symptoms. It is a novel viral infection that first emerged in Wuhan, China in December 2019 as a pneumonia of unknown etiology [1-4]. Coronaviruses infect humans and various animals, and subsequently cause respiratory, enteric, hepatic, and neurologic diseases [5,6]. Most human coronaviruses, apart from the SARS-CoV and Middle East Respiratory Syndrome (MERS)-CoV, infect the upper respiratory tract and lead to the development of mild respiratory symptoms with a low fatality rate. Conversely, the SARS-CoV and MERS-CoV mostly infect the lower respiratory tract and may lead to the development of severe respiratory symptoms, including shortness of breath and pneumonia.
- The novel SARS-CoV-2, the etiologic agent of COVID-19, is a single-stranded enveloped RNA virus of the family Coronaviridae that includes 4 genera: alphacoronaviruses, betacoronaviruses, deltacoronaviruses, and gammacoronaviruses. Of the human coronaviruses (HCoV), HCoV-229E and HCoV-NL3 are alphacoronaviruses, while HCoV-OC43 and HCoV-HKU1 are betacoronaviruses [3,4]. The SARS-CoV and the MERS-CoV, which first emerged in 2002 and 2012, respectively, belong to the betacoronavirus genera [1,7]. The SARS-CoV-2 is known to have a higher transmission rate and infectivity than SARS-CoV and MERS-CoV [7-9].
- Presence of a fever and a cough are the primary clinical manifestations of COVID-19 however, patients also exhibited other symptoms such as nausea, vomiting, diarrhea, and abdominal discomfort [3]. Presently, the most common method used for COVID-19 diagnosis is the detection of SARS-CoV-2 in upper and lower respiratory specimens, including nasopharyngeal swabs, oropharyngeal swabs, sputum, lower respiratory tract aspirates, and bronchoalveolar lavage. Genetic testing methods, such as real-time reverse transcription polymerase chain reaction (RT-PCR), are the standard methods of laboratory testing for COVID-19 that are currently in use in most countries.
- In the present study, we investigated whether SARS-CoV-2, which infects humans and may subsequently trigger the development of various clinical symptoms, can be detected in body fluids such as serum, urine, and stool, besides respiratory specimens. In addition, we aimed to isolate the virus from SARS-CoV-2-positive samples to determine viral infectivity.
Introduction
- 1. Specimens from COVID-19 patients
- To examine viral shedding in body parts other than the respiratory tract, and the infectivity of the detected virus, respiratory specimens such as nasopharyngeal swab, oropharyngeal swab or sputum, as well as serum, urine, and stool specimens were non-periodically sampled from 74 COVID-19 patients admitted in a hospital between January 19th and March 30th, 2020.
- Of the respiratory specimens, the upper respiratory samples were collected at least twice from all patients. Serum, urine, and stool samples were collected from 71, 54, and 38 patients, respectively. Each sample had been tested by the Korea Centers for Diseases Control and Prevention to monitor the SARS-CoV-2 infection status of the COVID-19 patients.
- 2. RNA extraction and real-time RT-PCR
- Serum samples were collected in a serum separation tube and centrifuged. Urine samples were centrifuged and the supernatants removed. The pellet was resuspended in 1-2 mL serum-free Dulbecco’s Modified Eagle Medium (DMEM). Each stool sample (1 g) was suspended in 10 mL of phosphate-buffered saline and was centrifuged to collect the supernatant for RNA extraction. RNA extraction and real-time RT-PCR was performed on the samples according to the method proposed by Kim et al in 2020 [10]. Briefly, RNA was extracted from 140 µL of the sample using a Qiagen viral RNA mini kit (Qiagen, Hilden, Germany) according to the method recommended by the manufacturer. Real-time RT-PCR was performed with the extracted RNA, and the cycle threshold value of the SARS-CoV-2 target gene was determined.
- 3. Virus isolation
- To isolate SARS-CoV-2 from samples that tested positive for the virus in real-time RT-PCR analyses, the samples were mixed with a 1:1 nystatin (10,000 U/mL) and penicillin-streptomycin (10,000 U/mL) mixture in a 1:4 ratio, and left to react at 4°C for 1 hour. The samples were then centrifuged at 400× g for 10 minutes and the supernatant was used as the inoculant. For the cell inoculation, cells were cultured from the CaCo-2 cell line (derived from human epithelial colorectal adenocarcinoma cells) in DMEM supplemented with 20% fetal bovine serum and 1% penicillin, and were incubated at 37°C, 5% CO2. On the day prior to inoculation, the cells were seeded at 2 × 105 cells/well into a 12-well plate. On the day of the primary inoculation, each well was replaced with DMEM supplemented with 2% fetal bovine serum, and 100 µL of the sample was used for the inoculation. The cells were cultured for 5 days and then harvested. The cell culture supernatant was centrifuged at 2,000 rpm for 10 minutes and the supernatant collected. The same method was repeated for a secondary inoculation. Five days after the secondary inoculation, the cell culture supernatant was harvested, centrifuged at 600 × g for 10 minutes, and the supernatant collected. To evaluate the viral replication process, RNA from the secondary inoculation cell culture supernatant was extracted and assessed for the presence of SARS-CoV-2 using real-time RT-PCR. The virus isolation experiment was conducted in a biosafety Level-3 facility.
- 4. Calculation of the viral copy number
- According to the standard curve, the plasmid DNA (E and RdRp genes) containing the linearized form of the PCR product (E gene, 113 bp; RdRp gene, 100 bp) were inserted into the pGEM-T Easy vector system by TA Cloning, followed by the transformation of DH10B cells (Promega Corporation, WI, USA), and the purification of plasmids using the QIAprep Spin Miniprep Kit and Qiagen Plasmid Plus Midikit (Qiagen, Hilden, Germany). The plasmid concentration was determined by measuring optical density with a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, USA), and the number of viral genome equivalent copies was calculated using the equation: amount of DNA (E gene: 7.4 × 1,011 copy/µL, RdRp gene: 8.4 × 1,011 copy/µL).
- 5. Ethics statement
- The study was approved by the Institutional Review Board at the Korea Centers for Disease Control and Prevention (2020-03-01-P-A). The board waived the requirement for written consent.
Materials and Methods
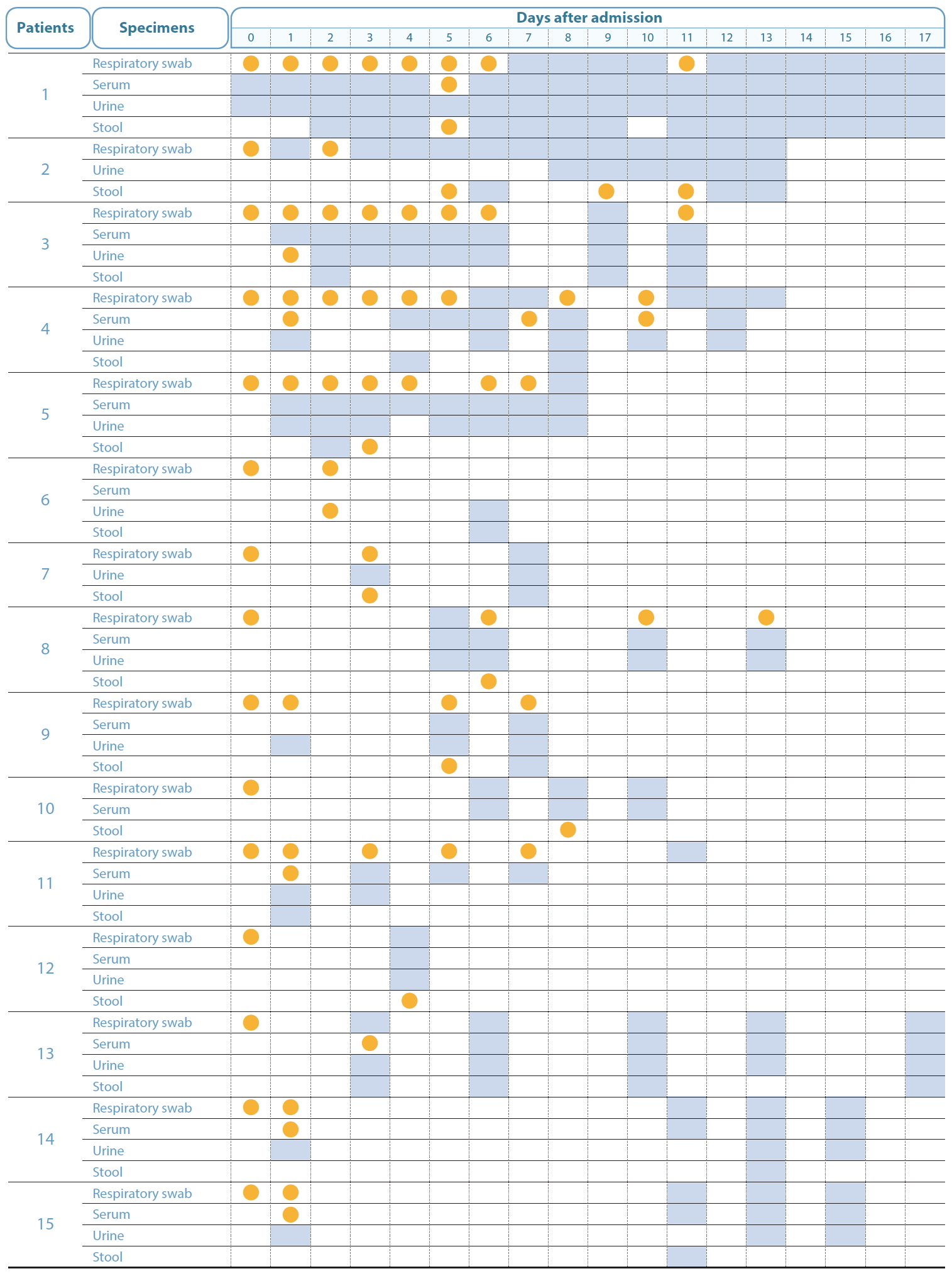
- The presence of SARS-CoV-2 was tested in serum, urine, and stool samples collected from 74 COVID-19 patients, using real-time RT-PCR. Of the patients, 44 were male and 30 were female aged 9-80 years (median 43 years). The maximum period of sample collection was 17 days from the date of disease confirmation. From the 74 patients, 323 serum samples, 247 urine samples, and 129 stool samples were collected and tested. The SARS-CoV-2 RNA detection rates were 2.8% (9/323), 0.8% (2/247), and 10.1% (13/129), respectively. In 15 patients (20.0%), SARS-CoV-2 was detected at least once in at least 1 of the 3 specimens, apart from the respiratory specimens (Figure 1). In 4 of these patients, the virus was detectable in the serum and stool samples for a longer period than in the respiratory samples. The virus was detected in serum samples of 6 patients, urine samples of 2 patients, and stool samples of 8 patients.
- The number of viral genome equivalent copies in the serum (19-6, 127 copies/µL), urine (49-109 copies/µL), and stool samples (27-27, 310 copies/µL), were calculated on the basis of the RdRp gene copy number. In 3 patients, the virus was not detected in the respiratory samples, although it was present in stool samples; in 2 of these patients, the virus could be detected in the stool samples at Days 7 and 9, even after it was no longer detectable in the respiratory samples. To isolate the virus, CaCo-2 cells were inoculated with the SARS-CoV-2 positive serum, urine, or stool sample. After primary inoculation, the cells were cultured for 5 days and then a secondary inoculation was performed however, the virus could not be isolated from the samples.
- There have been no published reports of SARS-CoV-2 viremia to date. The SARS-CoV-2 RNA level detected was 50% in plasma and 78% in serum samples at 7- or 14-days following hospitalization [11,12]. In other studies, the plasma viral load of SARS-CoV-2 peaked at 4-8 days after the disease onset, and followed a pattern of transient viremia, or occasionally persisted for more than 10 days, depending on the patient [13]. In this study, the viral RNA was detected in the serum of 5 patients at 3-6 days after onset. However, owing to irregular and discontinuous blood sampling, the duration for which the RNA was detectable could not be assessed (Figure 2). In addition, the inhibitory effect of the antiviral drug administered during treatment after hospitalization on viral replication cannot be excluded. Blood samples were collected 1-3 times within 6 days after onset in 46% of the patients. Since SARS-CoV-2 can be transmitted during the asymptomatic stage of infection before disease onset [14,15], future studies should investigate viremia during the asymptomatic stage and determine the rate of viremia.
- It has been reported that in SARS patients, viral shedding was detected in respiratory and stool specimens for up to 52 and 126 days, respectively [16]. However, in MERS patients, viral shedding has rarely been studied [17]. In a study on viral shedding in 37 MERS patients, the MERS-CoV RNA was detected in 93% of the lower respiratory samples for up to 3 weeks after disease confirmation, and viral RNA was detected in serum, stool, and urine samples [18]. However, virus could not be isolated from the stool or serum samples, which had the highest RNA concentration [19]. It has also been reported that MERS-CoV RNA was detected in stool samples, with the highest detection rate at Days 4-11 after onset, and was detected even at 30 days after onset in some patients [18,19].
- Although it has been reported that SARS-CoV-2 RNA was not detectable in the upper respiratory tract at 21 days on average after confirmation [14], there was a specific case wherein the duration of viral shedding was observed at 20 days and 37 days [20]. Apart from detection of SARS-CoV-2 RNA in the upper respiratory tract, it has been reported that SARS-CoV-2 RNA has been detected in stool samples. In a study on 28 COVID-19 patients, the most common clinical symptoms were a cough (28.6%), sore throat (28.6%), and fever (25.0%), while diarrhea (10.7%) was not a common symptom [3]. A study reported that 10% of the patients had diarrhea and experienced nausea 1-2 days before the onset of fever and respiratory symptoms however, there were no reports of viral RNA detection in the samples collected from these patients [18]. In another study, 11.6% of the hospitalized patients exhibited gastrointestinal symptoms at admittance and 49.5% exhibited such symptoms during their hospital stay, and depending on the presence of gastrointestinal symptoms, the viral RNA detection rates were 52.4% and 39.1%, respectively [21]. However, the study that reported the detection of SARS-CoV-2 RNA in stool samples did not investigate the infectivity of the detected viral RNA.
- The WHO recognizes the detection of SARS-CoV-2 RNA in lower and upper respiratory samples such as nasal swab, throat swab, and sputum as the official diagnostic method for COVID-19 testing. However, in addition to respiratory samples, viral RNA was also detected in serum, urine, or stool samples, suggesting the possibility of SARS-CoV-2 transmission via such specimens [22-25]. In this current study, SARS-CoV-2 RNA was detected in respiratory swabs as well as in serum, urine, and stool samples. Moreover, the virus from infected CaCo-2 cells was isolated to determine the infectivity of the viral RNA however, the presence of SARS-CoV-2 could not be confirmed in RNA-positive specimens, and the viral load was also considerably low. These results suggest that the SARS-CoV-2 is predominantly transmitted via the respiratory tract, and transmission via serum, urine, and stools is considerably limited.
Results and Discussion
-
Acknowledgements
- The authors thank all those who assisted in the collection and transport of patient samples. This study was funded by Korea Centers for Disease Control and Prevention (no.: 4800-4837-301).


- 1. Zhu N, Zhang D, Wang W, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 2020;382(8). 727−33.ArticlePubMedPMC
- 2. Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020;395:497−506.ArticlePubMedPMC
- 3. Kim ES, Chin BS, Kang CK, et al. Clinical course, and outcomes of patients with severe Acute respiratory syndrome coronavirus 2 infection: A preliminary report of the first 28 patients from the Korean cohort study on COVID-19. J Korean Med Sci 2020;35(13). e142.ArticlePubMedPMC
- 4. Jin Y, Yang H, Ji W, et al. Virology, epidemiology, pathogenesis, and control of COVID-19. Viruses 2020;12(4). E372.Article
- 5. Weiss SR, Leibowitz JL. Coronavirus pathogenesis. Adv Virus Res 2011;81:85−164.ArticlePubMedPMC
- 6. De Wilde AH, Snijder EJ, et al. Host factors in coronavirus replication. Curr Top Microbiol Immunol 2018;419:1−42.ArticlePubMed
- 7. Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020;579(7798). 270−3.ArticlePubMedPMCPDF
- 8. Li Q, Guan X, Wu P, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N Engl J Med 2020;382(13). 1199−207.ArticlePubMedPMC
- 9. Xu Z, Shi L, Wang Y, et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med 2020;8(4). 420−2.ArticlePubMedPMC
- 10. Kim JM, Chung YS, Jo JH, et al. Identification of coronavirus isolated from a patient in Korea with COVID-19. Osong Public Health Res Perspect 2020;11(1). 3−7.ArticlePubMedPMCPDF
- 11. Ng E, Hui D, Chan K, et al. Quantitative analysis, and prognostic implication of SARS coronavirus RNA in the plasma and serum of patients with severe acute respiratory syndrome. Clin Chem 2003;49(12). 1976−80.ArticlePubMedPMCPDF
- 12. Ng L, Wong M, Koh S, et al. Detection of severe acute respiratory syndrome coronavirus in blood of infected patients. J Clin Microbiol 2004;42(1). 347−50.ArticlePubMedPMC
- 13. Wang WK, Fang CT, Chen HL, et al. Detection of severe acute respiratory syndrome coronavirus RNA in plasma during the course of infection. J Clin Microbiol 2005;43(2). 962−5.ArticlePubMedPMC
- 14. Zou L, Ruan F, Huang M, et al. SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med 2020;382(12). 1177−9.ArticlePubMedPMC
- 15. He G, Sun W, Fang P, et al. The clinical feature of silent infections of novel coronavirus infection (COVID-19) in Wenzhou.[Epub ahead of print]. J Med Virol 2020;[Epub ahead of print]. Epub 2020 Apr 10.
- 16. Liu W, Tang F, Fontanet A, et al. Long-term SARS coronavirus excretion from patient cohort, China. Emerg Infect Dis 2004;10(10). 1841−3.ArticlePubMedPMC
- 17. Arabi YM, Arifi AA, Balkhy HH, et al. Clinical course and outcomes of critically ill patients with Middle East respiratory syndrome coronavirus infection. Ann Intern Med 2014;160(6). 389−97.ArticlePubMed
- 18. Corman VM, Albarrak AM, Omrani AS, et al. Viral shedding and antibody response in 37 patients with middle east respiratory syndrome coronavirus infection. Clin Infect Dis 2016;62(4). 477−83.ArticlePubMedPDF
- 19. Chan KH, Poon LL, Cheng VC, et al. Detection of SARS coronavirus in patients with suspected SARS. Emerg Infect Dis 2004;10(2). 294−9.ArticlePubMedPMC
- 20. Zhou F, Yu T, Du R, et al. Clinical course, and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020;395:1054−62.ArticlePubMedPMC
- 21. Lin L, Jiang X, Zhang Z, et al. Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut 2020;[Epub ahead of print]. Epub 2020 Apr 2.Article
- 22. Yeo C, Kaushal S, Yeo D. Enteric involvement of coronaviruses: Is faecal-oral transmission of SARS-CoV-2 possible? 2020;5(4). 335−7.Article
- 23. Chen Y, Chen L, Deng Q, et al. The presence of SARS-CoV-2 RNA in feces of COVID-19 patients. J Med Virol 2020;[Epub ahead of print]. Epub 2020 Apr 3.Article
- 24. Cao Y, Liu X, Xiong L, Cai K. Imaging, and clinical features of patients with 2019 novel coronavirus SARS-CoV-2: A systematic review and meta-analysis. J Med Virol 2020;J Med Virol 2020. [Epub ahead of print]. Epub 2020 Apr 3.Article
- 25. Tian Y, Rong L, Nian W, et al. Review article: Gastrointestinal features in COVID-19 and the possibility of faecal transmission. Aliment Pharmacol Ther 2020;51(9). 843−51.ArticlePubMedPMC
References
Figure & Data
References
Citations

- Natural SARS-CoV-2 infection in dogs: Determination of viral loads, distributions, localizations, and pathology
Chutchai Piewbang, Panida Poonsin, Pattiya Lohavicharn, Panitnan Punyathi, Sawang Kesdangsakonwut, Tanit Kasantikul, Somporn Techangamsuwan
Acta Tropica.2024; 249: 107070. CrossRef - Pharmacology of a Plant Virus Immunotherapy Candidate for Peritoneal Metastatic Ovarian Cancer
Anthony O. Omole, Jessica Fernanda Affonso de Oliveira, Lucas Sutorus, Nicole F. Steinmetz
ACS Pharmacology & Translational Science.2024; 7(2): 445. CrossRef - Optimal Selection of Sampling Points within Sewer Networks for Wastewater-Based Epidemiology Applications
Yao Yao, Yibo Zhu, Regina Nogueira, Frank Klawonn, Markus Wallner
Methods and Protocols.2024; 7(1): 6. CrossRef - Detection of SARS-CoV-2 and Omicron variant RNA in wastewater samples from Manila, Philippines
Jessamine Gail M. Inson, Bikash Malla, Divina M. Amalin, Thaddeus M. Carvajal, Ma. Luisa D. Enriquez, Soichiro Hirai, Sunayana Raya, Aulia Fajar Rahmani, Made Sandhyana Angga, Niva Sthapit, Sadhana Shrestha, Annisa Andarini Ruti, Tomoko Takeda, Masaaki Ki
Science of The Total Environment.2024; 919: 170921. CrossRef - Development of an Indexed Score to Identify the Most Suitable Biological Material to Assess SARS-CoV-2
Marina Almeida-Silva, Renata Cervantes, Edna Ribeiro, Ana Marques-Ramos
Applied Sciences.2024; 14(7): 2761. CrossRef - Norovirus and Coronavirus Risks in Food Service Settings: A Systematic Review for Future Research
Bendegul Okumus
Journal of Culinary Science & Technology.2023; 21(1): 71. CrossRef - Detection of the SARS-CoV-2 Genome in Peripheral Blood Mononuclear Cells of Hospitalized Patients With COVID-19 Infection
Mastaneh Alinezhadi, Maniya Arshadi, Mojtaba Rasti, Niloofar Neisi, Mehdi Parsanahad
Jundishapur Journal of Microbiology.2023;[Epub] CrossRef - Partial sequence conservation of SARS‐CoV‐2 NSP‐2, NSP‐12, and Spike in stool samples from Abadan, Iran
Milad Zandi, Saber Soltani, Alireza Tabibzadeh, Sepideh Nasimzadeh, Emad Behboudi, Armin Zakeri, Yousef Erfani, Shokrollah Salmanzadeh, Samaneh Abbasi
Biotechnology and Applied Biochemistry.2023; 70(1): 201. CrossRef - The Inhibition and Variability of Two Different RT-qPCR Assays Used for Quantifying SARS-CoV-2 RNA in Wastewater
George Scott, Nicholas Evens, Jonathan Porter, David I. Walker
Food and Environmental Virology.2023; 15(1): 71. CrossRef - COVID-19 and Glomerular Diseases
Nattawat Klomjit, Ladan Zand, Lynn D. Cornell, Mariam Priya Alexander
Kidney International Reports.2023; 8(6): 1137. CrossRef - Faecal shedding models for SARS-CoV-2 RNA among hospitalised patients and implications for wastewater-based epidemiology
Till Hoffmann, Justin Alsing
Journal of the Royal Statistical Society Series C:.2023; 72(2): 330. CrossRef - Investigation of SARS-CoV-2 using RT-PCR in vaginal swab samples of female patients with a diagnosis of severe COVID-19
Deniz Erdem, Bircan Kayaaslan, Esra Yakisik Cakir, Bedia Dinc, Dilek Asilturk, Fisun Kirca, Fatih Segmen, Isil Ozkocak Turan, Rahmet Guner
Taiwanese Journal of Obstetrics and Gynecology.2023; 62(2): 270. CrossRef - SARS-CoV-2 in the environment: Contamination routes, detection methods, persistence and removal in wastewater treatment plants
Ali Atoui, Christophe Cordevant, Thierry Chesnot, Benoît Gassilloud
Science of The Total Environment.2023; 881: 163453. CrossRef - Deciphering the Relationship between SARS-CoV-2 and Cancer
Michele Costanzo, Maria Anna Rachele De Giglio, Giovanni Nicola Roviello
International Journal of Molecular Sciences.2023; 24(9): 7803. CrossRef - Fecal-oral transmission of SARS-CoV-2: A systematic review of evidence from epidemiological and experimental studies
Martin Brink Termansen, Sebastian Frische
American Journal of Infection Control.2023; 51(12): 1430. CrossRef - Wastewater surveillance beyond COVID-19: a ranking system for communicable disease testing in the tri-county Detroit area, Michigan, USA
Zachary Gentry, Liang Zhao, Russell A. Faust, Randy E. David, John Norton, Irene Xagoraraki
Frontiers in Public Health.2023;[Epub] CrossRef - Clinical epidemiology of COVID-19 among hospitalized children in rural western Kenya
Adino Tesfahun Tsegaye, Christina Sherry, Chrisantus Oduol, Joyce Otieno, Doreen Rwigi, Mary Masheti, Irene Machura, Meshack Liru, Joyce Akuka, Deborah Omedo, Samwel Symekher, Samoel A. Khamadi, Lynda Isaaka, Morris Ogero, Livingstone Mumelo, James A. Ber
PLOS Global Public Health.2023; 3(6): e0002011. CrossRef - SARS-CoV-2 viral liver aggregates and scarce parenchymal infection implicate systemic disease as a driver of abnormal liver function
Brian J. Pepe-Mooney, Colton J. Smith, Marc S. Sherman, Trista E. North, Robert F. Padera, Wolfram Goessling
Hepatology Communications.2023;[Epub] CrossRef - Estimating the prevalence of COVID-19 cases through the analysis of SARS-CoV-2 RNA copies derived from wastewater samples from North Dakota
Bong-Jin Choi, Scott Hoselton, Grace N. Njau, I.G.C.G. Idamawatta, Paul Carson, John McEvoy
Global Epidemiology.2023; 6: 100124. CrossRef - SARS-CoV-2 Detection and COVID-19 Diagnosis: A Bird’s Eye View
Abdullah R. Alzahrani, Qaiser Jabeen, Imran Shahid, Saeed S. Al-Ghamdi, Naiyer Shahzad, Sidra Rehman, Alanood S. Algarni, Ghazi A. Bamagous, Ibrahim Mufadhi M. AlanazI, Ibrahim Abdel Aziz Ibrahim
Reviews on Recent Clinical Trials.2023; 18(3): 181. CrossRef - SARS-CoV-2 excretion kinetics in nasopharyngeal and stool samples from the pediatric population
Haifa Khemiri, Mariem Gdoura, Samar Ben Halima, Henda Krichen, Cesare Cammà, Alessio Lorusso, Massimo Ancora, Adriano Di Pasquale, Asma Cherni, Henda Touzi, Amel Sadraoui, Zina Meddeb, Nahed Hogga, Radhia Ammi, Henda Triki, Sondes Haddad-Boubaker
Frontiers in Medicine.2023;[Epub] CrossRef - EN PRENSA. SARS-COV-2 RNA decay on raw environmental aquatic matrices. EN PRENSA
Juan Daniel Lira-Morales, Jose Andrés Medrano-Felix, Celida Isabel Martinez-Rodriguez, Nohelia Castro-del Campo, Cristobal Chaidez-Quiroz
Revista Bio Ciencias.2023;[Epub] CrossRef - Features of SARS-CoV-2 Replication in Various Types of Reptilian and Fish Cell Cultures
Yulia Kononova, Lyubov Adamenko, Evgeniya Kazachkova, Mariya Solomatina, Svetlana Romanenko, Anastasia Proskuryakova, Yaroslav Utkin, Marina Gulyaeva, Anastasia Spirina, Elena Kazachinskaia, Natalia Palyanova, Oksana Mishchenko, Alexander Chepurnov, Alexa
Viruses.2023; 15(12): 2350. CrossRef - New Insights into the Link between SARS-CoV-2 Infection and Renal Cancer
Vittoria Rago, Sabrina Bossio, Danilo Lofaro, Anna Perri, Silvia Di Agostino
Life.2023; 14(1): 52. CrossRef - Occurrence of SARS-CoV-2 in excreta, sewage, and environment: epidemiological significance and potential risks
Omar Amahmid, Youssef El Guamri, Youness Rakibi, Saadia Ouizat, Mohamed Yazidi, Bouchra Razoki, Khadija Kaid Rassou, Souad Asmama, Khadija Bouhoum, Driss Belghyti
International Journal of Environmental Health Rese.2022; 32(8): 1686. CrossRef - Twelve Months with COVID-19: What Gastroenterologists Need to Know
Giulia Concas, Michele Barone, Ruggiero Francavilla, Fernanda Cristofori, Vanessa Nadia Dargenio, Rossella Giorgio, Costantino Dargenio, Vassilios Fanos, Maria Antonietta Marcialis
Digestive Diseases and Sciences.2022; 67(7): 2771. CrossRef - SARS-CoV-2 loads in urine, sera and stool specimens in association with clinical features of COVID-19 patients
Déborah Anjos, Fabiola Souza Fiaccadori, Carolina do Prado Servian, Simone Gonçalves da Fonseca, Adriana Oliveira Guilarde, Moara Alves Santa Bárbara Borges, Fernanda Craveiro Franco, Bergmann Morais Ribeiro, Menira Souza
Journal of Clinical Virology Plus.2022; 2(1): 100059. CrossRef - Absence of virological and epidemiological evidence that SARS-CoV-2 poses COVID-19 risks from environmental fecal waste, wastewater and water exposures
Mark D. Sobsey
Journal of Water and Health.2022; 20(1): 126. CrossRef - Discrimination and surveillance of infectious severe acute respiratory syndrome Coronavirus 2 in wastewater using cell culture and RT-qPCR
Silvia Monteiro, Daniela Rente, Mónica V. Cunha, Tiago A. Marques, Eugénia Cardoso, João Vilaça, Norberta Coelho, Nuno Brôco, Marta Carvalho, Ricardo Santos
Science of The Total Environment.2022; 815: 152914. CrossRef - A collaborative robotic solution to partly automate SARS-CoV-2 serological tests in small facilities
Andrea Maria Zanchettin, Federica Facciotti
SLAS Technology.2022; 27(1): 100. CrossRef - The Impact of COVID-19 on Blood Transfusion Services: A Systematic Review and Meta-Analysis
Cindy Chiem, Khader Alghamdi, Thao Nguyen, Joon Hee Han, Hongzhao Huo, Denise Jackson
Transfusion Medicine and Hemotherapy.2022; 49(2): 107. CrossRef - Development of an in vitro model for animal species susceptibility to SARS-CoV-2 replication based on expression of ACE2 and TMPRSS2 in avian cells
Darrell R. Kapczynski, Ryan Sweeney, Erica Spackman, Mary Pantin-Jackwood, David L. Suarez
Virology.2022; 569: 1. CrossRef - Kidney Injury in COVID-19: Epidemiology, Molecular Mechanisms and Potential Therapeutic Targets
J. Pedro Teixeira, Sharon Barone, Kamyar Zahedi, Manoocher Soleimani
International Journal of Molecular Sciences.2022; 23(4): 2242. CrossRef - SARS-CoV-2 shedding sources in wastewater and implications for wastewater-based epidemiology
Xuan Li, Jagadeeshkumar Kulandaivelu, Ying Guo, Shuxin Zhang, Jiahua Shi, Jake O’Brien, Sudipti Arora, Manish Kumar, Samendra P. Sherchan, Ryo Honda, Greg Jackson, Stephen P. Luby, Guangming Jiang
Journal of Hazardous Materials.2022; 432: 128667. CrossRef - An Ensemble Learning Model for COVID-19 Detection from Blood Test Samples
Olusola O. Abayomi-Alli, Robertas Damaševičius, Rytis Maskeliūnas, Sanjay Misra
Sensors.2022; 22(6): 2224. CrossRef - Correlation of SARS‑CoV‑2 to cancer: Carcinogenic or anticancer? (Review)
Ying-Shuang Li, Hua-Cheng Ren, Jian-Hua Cao
International Journal of Oncology.2022;[Epub] CrossRef - Prolonged fecal shedding of SARS‐CoV‐2 in a young immunocompetent COVID‐19 patient: A case report and literature overview
Fang Xiao, Peiqi Wan, Qiuling Wei, Guiyang Wei, Yanhong Yu
Journal of Medical Virology.2022; 94(7): 3133. CrossRef - Lead time of early warning by wastewater surveillance for COVID-19: Geographical variations and impacting factors
Manish Kumar, Guangming Jiang, Alok Kumar Thakur, Shreya Chatterjee, Tanushree Bhattacharya, Sanjeeb Mohapatra, Tushara Chaminda, Vinay Kumar Tyagi, Meththika Vithanage, Prosun Bhattacharya, Long D. Nghiem, Dibyendu Sarkar, Christian Sonne, Jürgen Mahlkne
Chemical Engineering Journal.2022; 441: 135936. CrossRef - Possible Cross-Reactivity of Feline and White-Tailed Deer Antibodies against the SARS-CoV-2 Receptor Binding Domain
Trevor J. Hancock, Peyton Hickman, Niloo Kazerooni, Melissa Kennedy, Stephen A. Kania, Michelle Dennis, Nicole Szafranski, Richard Gerhold, Chunlei Su, Tom Masi, Stephen Smith, Tim E. Sparer, Tom Gallagher
Journal of Virology.2022;[Epub] CrossRef - Artificial neural network-based estimation of COVID-19 case numbers and effective reproduction rate using wastewater-based epidemiology
Guangming Jiang, Jiangping Wu, Jennifer Weidhaas, Xuan Li, Yan Chen, Jochen Mueller, Jiaying Li, Manish Kumar, Xu Zhou, Sudipti Arora, Eiji Haramoto, Samendra Sherchan, Gorka Orive, Unax Lertxundi, Ryo Honda, Masaaki Kitajima, Greg Jackson
Water Research.2022; 218: 118451. CrossRef - SARS-CoV-2 Viral Load in the Pulmonary Compartment of Critically Ill COVID-19 Patients Correlates with Viral Serum Load and Fatal Outcomes
Mario Ynga-Durand, Henrike Maaß, Marko Milošević, Fran Krstanović, Marina Pribanić Matešić, Stipan Jonjić, Alen Protić, Ilija Brizić, Alan Šustić, Luka Čičin-Šain
Viruses.2022; 14(6): 1292. CrossRef - Application of human RNase P normalization for the realistic estimation of SARS-CoV-2 viral load in wastewater: A perspective from Qatar wastewater surveillance
Shimaa S. El-Malah, Jayaprakash Saththasivam, Khadeeja Abdul Jabbar, Arun K.K., Tricia A. Gomez, Ayeda A. Ahmed, Yasmin A. Mohamoud, Joel A. Malek, Laith J. Abu Raddad, Hussein A. Abu Halaweh, Roberto Bertollini, Jenny Lawler, Khaled A. Mahmoud
Environmental Technology & Innovation.2022; 27: 102775. CrossRef - Predictors of intensive care unit admission and mortality in SARS-CoV-2 infection: A cross sectional study at a tertiary care hospital
Ahmed Alkhamis, Yousef Alshamali, Wassim Chehadeh, Amar Jasem, Abdulaziz Alsayed Omar, Mohammad Alghounaim, Hussein Elsaaran, Sarah Al-Youha, Sulaiman Almazeedi, Moh A. Alkhamis, Salman Alsabah
Annals of Medicine and Surgery.2022; 80: 104097. CrossRef - Vertical outbreak of COVID-19 in high-rise buildings: The role of sewer stacks and prevention measures
Ying Guo, Xuan Li, Stephen Luby, Guangming Jiang
Current Opinion in Environmental Science & Health.2022; 29: 100379. CrossRef - Current trends in COVID-19 diagnosis and its new variants in physiological fluids: Surface antigens, antibodies, nucleic acids, and RNA sequencing
Menna Mostafa, Ahmed Barhoum, Ekin Sehit, Hossam Gewaid, Eslam Mostafa, Mohamed M. Omran, Mohga S. Abdalla, Fatehy M. Abdel-Haleem, Zeynep Altintas, Robert J. Forster
TrAC Trends in Analytical Chemistry.2022; 157: 116750. CrossRef - The Persistence Time of SARS-CoV-2 RNA in Hospitalized COVID-19
Patients: A Prospective Study
Jila Yavarian, Mohammad Javaherian, Somayeh Shatizadeh Malekshahi, Zahra Ahmadinejad, Fatemeh Ghiasvand, Elaheh Ahmadi, Ladan Abbasian, Malihe Hasannezhad, Narges Dabaghipour, Nazanin-Zahra Shafiei-Jandaghi, Talat Mokhtari-Azad, Mahboubeh Hajiabdolbaghi
Infectious Disorders - Drug Targets.2022;[Epub] CrossRef - Evaluation of the presence of SARS-CoV-2 in vaginal and anal swabs of women with omicron variants of SARS-CoV-2 infection
Ding Liu, Yunfu Zhang, Dongfeng Chen, Xianhua Wang, Fuling Huang, Ling Long, Xiuhui Zheng
Frontiers in Microbiology.2022;[Epub] CrossRef - Immune-profiling of SARS-CoV-2 viremic patients reveals dysregulated innate immune responses
Xiaoming Sun, Ce Gao, Ke Zhao, Yanhui Yang, Yelizaveta Rassadkina, Jesse Fajnzylber, James Regan, Jonathan Z. Li, Mathias Lichterfeld, Xu G. Yu
Frontiers in Immunology.2022;[Epub] CrossRef - Pathogens inactivation in nutrient recovery from urine: A review
Kangning Xu, Jun Lu, Lanlan Hu, Jiyun Li, Shikun Cheng, Min Zheng, Chengwen Wang
Frontiers in Environmental Science.2022;[Epub] CrossRef - Sars-Cov-2 Shedding Sources in Wastewater and Implications for Wastewater-Based Epidemiology
Xuan Li, Jagadeeshkumar Kulandaivelu, Ying Guo, Shuxin Zhang, Jiahua Shi, Jake O’Brien, Sudipti Arora, Manish Kumar, Samendra P. Sherchan, Ryo Honda, Greg Jackson, Stephen Luby, Guangming Jiang
SSRN Electronic Journal .2022;[Epub] CrossRef - Severe acute respiratory syndrome coronavirus‐2: implications for blood safety and sufficiency
Philip Kiely, Veronica C. Hoad, Clive R. Seed, Iain B. Gosbell
Vox Sanguinis.2021; 116(2): 155. CrossRef - Is recurrence possible in coronavirus disease 2019 (COVID-19)? Case series and systematic review of literature
Anna Gidari, Marco Nofri, Luca Saccarelli, Sabrina Bastianelli, Samuele Sabbatini, Silvia Bozza, Barbara Camilloni, Igino Fusco-Moffa, Claudia Monari, Edoardo De Robertis, Antonella Mencacci, Daniela Francisci
European Journal of Clinical Microbiology & Infect.2021; 40(1): 1. CrossRef - Hunting coronavirus by transmission electron microscopy – a guide to SARS‐CoV‐2‐associated ultrastructural pathology in COVID‐19 tissues
Helmut Hopfer, Martin C Herzig, Rainer Gosert, Thomas Menter, Jürgen Hench, Alexandar Tzankov, Hans H Hirsch, Sara E Miller
Histopathology.2021; 78(3): 358. CrossRef - CRISPR-based biosensing systems: a way to rapidly diagnose COVID-19
Majid Vatankhah, Amir Azizi, Anahita Sanajouyan Langeroudi, Sajad Ataei Azimi, Imaneh Khorsand, Mohammad Amin Kerachian, Jamshid Motaei
Critical Reviews in Clinical Laboratory Sciences.2021; 58(4): 225. CrossRef - The virus that shook the world: questions and answers about SARS-CoV-2 and COVID-19
Radostina Alexandrova, Pencho Beykov, Dobrin Vassilev, Marko Jukić, Črtomir Podlipnik
Biotechnology & Biotechnological Equipment.2021; 35(1): 74. CrossRef - Comprehensive analysis of COVID-19 during pregnancy
Kathryn M. Moore, Mehul S. Suthar
Biochemical and Biophysical Research Communication.2021; 538: 180. CrossRef - Innate Immunity Plays a Key Role in Controlling Viral Load in COVID-19: Mechanistic Insights from a Whole-Body Infection Dynamics Model
Prashant Dogra, Javier Ruiz-Ramírez, Kavya Sinha, Joseph D. Butner, Maria J. Peláez, Manmeet Rawat, Venkata K. Yellepeddi, Renata Pasqualini, Wadih Arap, H. Dirk Sostman, Vittorio Cristini, Zhihui Wang
ACS Pharmacology & Translational Science.2021; 4(1): 248. CrossRef - Transmission of SARS-CoV-2: A Review of Viral, Host, and Environmental Factors
Eric A. Meyerowitz, Aaron Richterman, Rajesh T. Gandhi, Paul E. Sax
Annals of Internal Medicine.2021; 174(1): 69. CrossRef - CRISPR-based detection of SARS-CoV-2: A review from sample to result
Reza Nouri, Zifan Tang, Ming Dong, Tianyi Liu, Aneesh Kshirsagar, Weihua Guan
Biosensors and Bioelectronics.2021; 178: 113012. CrossRef - Evaluation of two fluorescence immunoassays for the rapid detection of SARS-CoV-2 antigen—new tool to detect infective COVID-19 patients
Lorena Porte, Paulette Legarraga, Mirentxu Iruretagoyena, Valeska Vollrath, Gabriel Pizarro, Jose Munita, Rafael Araos, Thomas Weitzel
PeerJ.2021; 9: e10801. CrossRef - Rapid detection of SARS-CoV-2, replicating or non-replicating, using RT-PCR
Ming Liao, Jianmin Wu, Manman Dai, Huanan Li, Nan Yan, Runyu Yuan, Chungen Pan
International Journal of Infectious Diseases.2021; 104: 471. CrossRef - A review on human body fluids for the diagnosis of viral infections: scope for rapid detection of COVID-19
Sphurti S Adigal, Nidheesh V Rayaroth, Reena V John, Keerthilatha M Pai, Sulatha Bhandari, Aswini Kumar Mohapatra, Jijo Lukose, Ajeetkumar Patil, Aseefhali Bankapur, Santhosh Chidangil
Expert Review of Molecular Diagnostics.2021; 21(1): 31. CrossRef - COVID-19 (SARS-CoV-2) outbreak monitoring using wastewater-based epidemiology in Qatar
Jayaprakash Saththasivam, Shimaa S. El-Malah, Tricia A. Gomez, Khadeeja A. Jabbar, Reshma Remanan, Arun K. Krishnankutty, Oluwaseun Ogunbiyi, Kashif Rasool, Sahel Ashhab, Sergey Rashkeev, Meryem Bensaad, Ayeda A. Ahmed, Yasmin A. Mohamoud, Joel A. Malek,
Science of The Total Environment.2021; 774: 145608. CrossRef - A critical review on SARS-CoV-2 infectivity in water and wastewater. What do we know?
Alexandre Giacobbo, Marco Antônio Siqueira Rodrigues, Jane Zoppas Ferreira, Andréa Moura Bernardes, Maria Norberta de Pinho
Science of The Total Environment.2021; 774: 145721. CrossRef - Uncertainties in estimating SARS-CoV-2 prevalence by wastewater-based epidemiology
Xuan Li, Shuxin Zhang, Jiahua Shi, Stephen P. Luby, Guangming Jiang
Chemical Engineering Journal.2021; 415: 129039. CrossRef - Re-estimation of basic reproduction number of COVID-19 based on the epidemic curve by symptom onset date
K. Hong, S. J. Yum, J. H. Kim, B. C. Chun
Epidemiology and Infection.2021;[Epub] CrossRef - Implications of inadequate water and sanitation infrastructure for community spread of COVID-19 in remote Alaskan communities
Laura Eichelberger, Subhabrata Dev, Tricia Howe, David L. Barnes, Eric Bortz, Brandon R. Briggs, Patricia Cochran, Aaron D. Dotson, Devin M. Drown, Micah B. Hahn, Kaitlin Mattos, Srijan Aggarwal
Science of The Total Environment.2021; 776: 145842. CrossRef - Validation of a new automated chemiluminescent anti-SARS-CoV-2 IgM and IgG antibody assay system detecting both N and S proteins in Japan
Rin Yokoyama, Makoto Kurano, Yoshifumi Morita, Takuya Shimura, Yuki Nakano, Chungen Qian, Fuzhen Xia, Fan He, Yoshiro Kishi, Jun Okada, Naoyuki Yoshikawa, Yutaka Nagura, Hitoshi Okazaki, Kyoji Moriya, Yasuyuki Seto, Tatsuhiko Kodama, Yutaka Yatomi, Kateri
PLOS ONE.2021; 16(3): e0247711. CrossRef - Sewage as a Possible Transmission Vehicle During a Coronavirus Disease 2019 Outbreak in a Densely Populated Community: Guangzhou, China, April 2020
Jun Yuan, Zongqiu Chen, Chenghua Gong, Hui Liu, Baisheng Li, Kuibiao Li, Xi Chen, Conghui Xu, Qinlong Jing, Guocong Liu, Pengzhe Qin, Yufei Liu, Yi Zhong, Lijuan Huang, Bao-Ping Zhu, Zhicong Yang
Clinical Infectious Diseases.2021; 73(7): e1795. CrossRef - Pathogenesis of coronavirus disease 2019-associated kidney injury
Kelly D. Smith, Shreeram Akilesh
Current Opinion in Nephrology & Hypertension.2021; 30(3): 324. CrossRef - Rapid and unamplified identification of COVID-19 with morpholino-modified graphene field-effect transistor nanosensor
Jiahao Li, Ding Wu, Yi Yu, Tingxian Li, Kun Li, Meng-Meng Xiao, Yirong Li, Zhi-Yong Zhang, Guo-Jun Zhang
Biosensors and Bioelectronics.2021; 183: 113206. CrossRef - Focused role of nanoparticles against COVID-19: Diagnosis and treatment
Mohammed Ali Dheyab, Pegah Moradi Khaniabadi, Azlan Abdul Aziz, Mahmood S. Jameel, Baharak Mehrdel, Ammar A. Oglat, Hawraa Ali Khaleel
Photodiagnosis and Photodynamic Therapy.2021; 34: 102287. CrossRef - Determining the communicable period of SARS-CoV-2: A rapid review of the literature, March to September 2020
Mina Park, Colleen Pawliuk, Tribesty Nguyen, Amanda Griffitt, Linda Dix-Cooper, Nadia Fourik, Martin Dawes
Eurosurveillance.2021;[Epub] CrossRef - Maternal respiratory SARS-CoV-2 infection in pregnancy is associated with a robust inflammatory response at the maternal-fetal interface
Alice Lu-Culligan, Arun R. Chavan, Pavithra Vijayakumar, Lina Irshaid, Edward M. Courchaine, Kristin M. Milano, Zhonghua Tang, Scott D. Pope, Eric Song, Chantal B.F. Vogels, William J. Lu-Culligan, Katherine H. Campbell, Arnau Casanovas-Massana, Santos Be
Med.2021; 2(5): 591. CrossRef - SARS-CoV-2: Pathogenesis, Molecular Targets and Experimental Models
G. Kanimozhi, B. Pradhapsingh, Charan Singh Pawar, Haseeb A. Khan, Salman H. Alrokayan, N. Rajendra Prasad
Frontiers in Pharmacology.2021;[Epub] CrossRef - Duration of culturable SARS-CoV-2 within different specimens among mild and severe COVID-19 patients: A longitudinal study
Yae Jee Baek, Youn-Jung Lee, Jae Sun Yoon, Yujin Sohn, Yunsuk Cho, Moo Hyun Kim, Jong Hoon Hyun, Eun Hwa Lee, Ki Hyun Lee, Jung Ho Kim, Jin Young Ahn, Su Jin Jeong, Nam Su Ku, Joon-Sup Yeom, Jun Yong Choi
Journal of Infection.2021; 83(1): e29. CrossRef - Tools and Techniques for Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2)/COVID-19 Detection
Seyed Hamid Safiabadi Tali, Jason J. LeBlanc, Zubi Sadiq, Oyejide Damilola Oyewunmi, Carolina Camargo, Bahareh Nikpour, Narges Armanfard, Selena M. Sagan, Sana Jahanshahi-Anbuhi
Clinical Microbiology Reviews.2021;[Epub] CrossRef - The risk of transmitting the coronavirus to the perioperative team through aerosols produced in the operating room bathrooms
Mahdieh Soleimani, Amirmohammad Merajikhah, Atefeh Beigi-khoozani
Perioperative Care and Operating Room Management.2021; 24: 100179. CrossRef - Monitoring the presence and persistence of SARS-CoV-2 in water-food-environmental compartments: State of the knowledge and research needs
Bashir Adelodun, Fidelis Odedishemi Ajibade, AbdulGafar Olatunji Tiamiyu, Nathaniel Azubuike Nwogwu, Rahmat Gbemisola Ibrahim, Pankaj Kumar, Vinod Kumar, Golden Odey, Krishna Kumar Yadav, Afzal Husain Khan, Marina M.S. Cabral-Pinto, Kola Yusuff Kareem, Ha
Environmental Research.2021; 200: 111373. CrossRef - Nucleic Acid Testing of SARS-CoV-2
Hee Min Yoo, Il-Hwan Kim, Seil Kim
International Journal of Molecular Sciences.2021; 22(11): 6150. CrossRef - What’s in the Pipeline? Evidence on the Transmission of SARS-CoV-2 via Building Wastewater Plumbing Systems
Thomas Dight, Michael Gormley
Frontiers in Built Environment.2021;[Epub] CrossRef - Kidney transplantation from a SARS‐CoV‐2‐positive donor for the recipients with immunity after COVID‐19
Lina Puodziukaite, Mindaugas Serpytis, Aurelija Kundrotaite, Jurate Sipylaite, Marius Miglinas, Marta Monika Janusaite, Egle Asakiene, Erika Milisauskaite, Arunas Zelvys, Oleg Aliancevic, Ligita Jancoriene
Transplant Infectious Disease.2021;[Epub] CrossRef - Evidence of SARS-CoV-2 Infection in Cells, Tissues, and Organs and the Risk of Transmission Through Transplantation
Amaury Gaussen, Laura Hornby, Gary Rockl, Sheila O’Brien, Gilles Delage, Ruth Sapir-Pichhadze, Steven J. Drews, Matthew J. Weiss, Antoine Lewin
Transplantation.2021; 105(7): 1405. CrossRef - Evaluation of SARS-CoV-2 in the Vaginal Secretions of Women with COVID-19: A Prospective Study
Elad Barber, Michal Kovo, Sophia Leytes, Ron Sagiv, Eran Weiner, Orna Schwartz, Margarita Mashavi, Keren Holtzman, Jacob Bar, Anat Engel, Shimon Ginath
Journal of Clinical Medicine.2021; 10(12): 2735. CrossRef - Detection of SARS-CoV-2 RNA in serum is associated with increased mortality risk in hospitalized COVID-19 patients
Diego A. Rodríguez-Serrano, Emilia Roy-Vallejo, Nelly D. Zurita Cruz, Alexandra Martín Ramírez, Sebastián C. Rodríguez-García, Nuria Arevalillo-Fernández, José María Galván-Román, Leticia Fontán García-Rodrigo, Lorena Vega-Piris, Marta Chicot Llano, David
Scientific Reports.2021;[Epub] CrossRef - Needle Stick Injury From a COVID-19 Patient—Fear It or Forget It?
Vishakh C. Keri, Parul Kodan, Anubhav Gupta, Pankaj Jorwal
Journal of Bioethical Inquiry.2021; 18(3): 377. CrossRef - Detection of three pandemic causing coronaviruses from non-respiratory samples: systematic review and meta-analysis
Chandan Mishra, Suneeta Meena, Jitendra Kumar Meena, Suman Tiwari, Purva Mathur
Scientific Reports.2021;[Epub] CrossRef - Sewage Systems Surveillance for SARS-CoV-2: Identification of Knowledge Gaps, Emerging Threats, and Future Research Needs
Fatemeh Amereh, Masoud Negahban-Azar, Siavash Isazadeh, Hossein Dabiri, Najmeh Masihi, Mahsa Jahangiri-rad, Mohammad Rafiee
Pathogens.2021; 10(8): 946. CrossRef - Probiotics and Coronavirus disease 2019: think about the link
Suresh Kumar Angurana, Arun Bansal
British Journal of Nutrition.2021; 126(10): 1564. CrossRef - Role of Nucleocapsid Protein Antigen Detection for Safe End of Isolation of SARS-CoV-2 Infected Patients with Long Persistence of Viral RNA in Respiratory Samples
Antonella Mencacci, Alessio Gili, Anna Gidari, Elisabetta Schiaroli, Carla Russo, Elio Cenci, Barbara Camilloni, Alessandro Graziani, Arduino Melelli-Roia, Daniela Francisci, Fabrizio Stracci
Journal of Clinical Medicine.2021; 10(18): 4037. CrossRef - Designing Short Peptides to Block the Interaction of SARS-CoV-2 and Human ACE2 for COVID-19 Therapeutics
Abdul Basit, Asad Mustafa Karim, Muhammad Asif, Tanveer Ali, Jung Hun Lee, Jeong Ho Jeon, Shafiq ur Rehman, Sang Hee Lee
Frontiers in Pharmacology.2021;[Epub] CrossRef - Tracking SARS-CoV-2: Novel Trends and Diagnostic Strategies
Linda P. Guaman-Bautista, Erick Moreta-Urbano, Claudia G. Oña-Arias, Marbel Torres-Arias, Nikolaos C. Kyriakidis, Koray Malcı, Nestor Jonguitud-Borrego, Leonardo Rios-Solis, Espiridion Ramos-Martinez, Andrés López-Cortés, Carlos Barba-Ostria
Diagnostics.2021; 11(11): 1981. CrossRef - Probable Case of Vertical Transmission of SARS-CoV-2 in a Newborn in Mexico
Marcela Mendoza-Hernández, Ivette Huerta-Niño de Rivera, María Yoldi-Negrete, Pamela Saviñon-Tejeda, Rafael Franco-Cendejas, Luis Esaú López-Jácome, Iñaki Navarro-Castellanos
Neonatology.2021; 118(3): 364. CrossRef - Validation of a novel molecular assay to the diagnostic of COVID-19 based on real time PCR with high resolution melting
Beatriz Iandra da Silva Ferreira, Natália Lins da Silva-Gomes, Wagner Luis da Costa Nunes Pimentel Coelho, Vanessa Duarte da Costa, Vanessa Cristine de Souza Carneiro, Rafael Lopes Kader, Marisa Pimentel Amaro, Lívia Melo Villar, Fábio Miyajima, Soniza Vi
PLOS ONE.2021; 16(11): e0260087. CrossRef - Cross-validation of SARS-CoV-2 responses in kidney organoids and clinical populations
Louisa Helms, Silvia Marchiano, Ian B. Stanaway, Tien-Ying Hsiang, Benjamin A. Juliar, Shally Saini, Yan Ting Zhao, Akshita Khanna, Rajasree Menon, Fadhl Alakwaa, Carmen Mikacenic, Eric D. Morrell, Mark M. Wurfel, Matthias Kretzler, Jennifer L. Harder, Ch
JCI Insight.2021;[Epub] CrossRef - COVID-19 and Kidney Involvement - A Systematic Review
Guang Yang, Yang Liu, Jiebin Hou, Qingli Cheng
Integrative Medicine in Nephrology and Andrology.2021; 8(1): 4. CrossRef - Viable SARS-CoV-2 in various specimens from COVID-19 patients
Hye Won Jeong, Se-Mi Kim, Hee-Sung Kim, Young-Il Kim, Jun Hyoung Kim, Jun Yeon Cho, Sun-hyung Kim, Hyeran Kang, Seong-Gyu Kim, Su-Jin Park, Eun-Ha Kim, Young Ki Choi
Clinical Microbiology and Infection.2020; 26(11): 1520. CrossRef - Risk of transmission of severe acute respiratory syndrome coronavirus 2 by transfusion: A literature review
Jean‐François Leblanc, Marc Germain, Gilles Delage, Sheila OʼBrien, Steven J. Drews, Antoine Lewin
Transfusion.2020; 60(12): 3046. CrossRef - Potential sources, modes of transmission and effectiveness of prevention measures against SARS-CoV-2
G. Kampf, Y. Brüggemann, H.E.J. Kaba, J. Steinmann, S. Pfaender, S. Scheithauer, E. Steinmann
Journal of Hospital Infection.2020; 106(4): 678. CrossRef - Does SARS-CoV-2 Infect the Kidney?
Shaza Khan, Lihe Chen, Chin-Rang Yang, Viswanathan Raghuram, Syed J. Khundmiri, Mark A. Knepper
Journal of the American Society of Nephrology.2020; 31(12): 2746. CrossRef - Les Coronavirus humains
Michel Segondy
Revue Francophone des Laboratoires.2020; 2020(526): 32. CrossRef - In reply: an example of the evolution of evidence in a new disease
Benjamin Condon, Thomas Whish-Wilson, Niall F. Davis, Nathan Lawrentschuk
Future Oncology.2020; 16(35): 2903. CrossRef - Wastewater-Based Epidemiology (WBE) and Viral Detection in Polluted Surface Water: A Valuable Tool for COVID-19 Surveillance—A Brief Review
Maria de Lourdes Aguiar-Oliveira, Aline Campos, Aline R. Matos, Caroline Rigotto, Adriana Sotero-Martins, Paulo F. P. Teixeira, Marilda M. Siqueira
International Journal of Environmental Research an.2020; 17(24): 9251. CrossRef - Cave canem: urine is not urine in corona times
Kunz Yannic, Horninger Wolfgang, Pinggera Germar-Michael
Future Oncology.2020; 16(35): 2899. CrossRef - A Public Platform for Virtual IoT-Based Monitoring and Tracking of COVID-19
Younchan Jung, Ronnel Agulto
Electronics.2020; 10(1): 12. CrossRef


 PubReader
PubReader ePub Link
ePub Link Cite
Cite